Maček lab
Post-translational regulation in the autotrophy-to-heterotrophy switch in Synechocystis
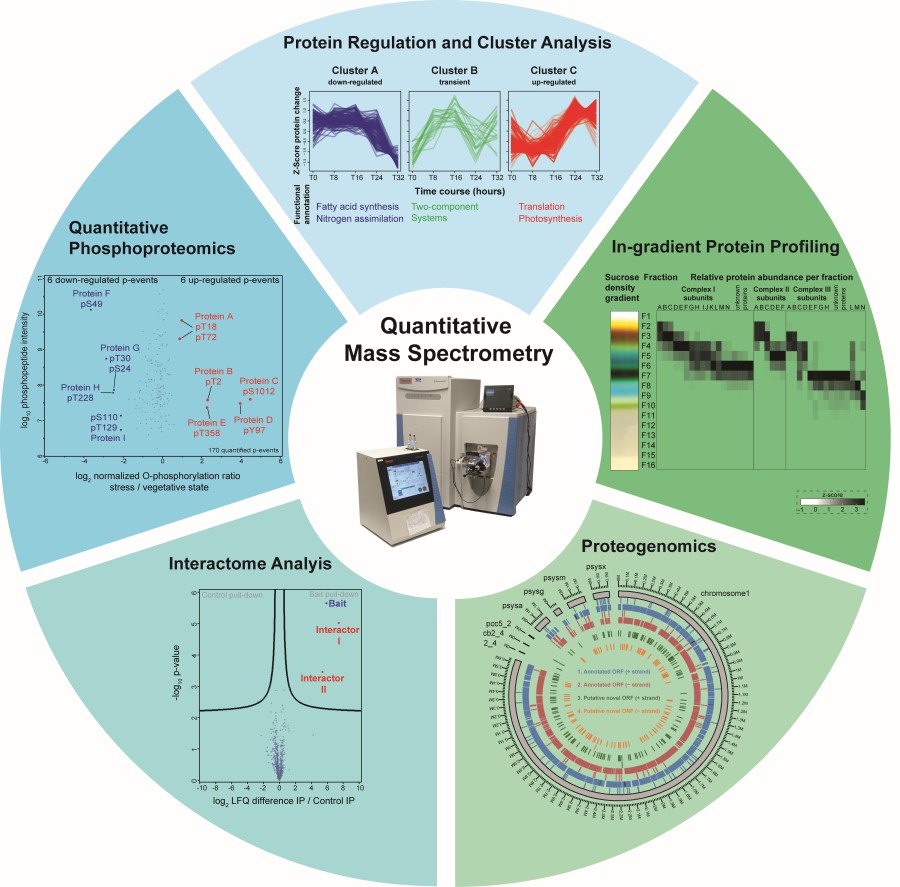
Protein phosphorylation on Ser/Thr and Tyr residues has been identified in Synechocystis sp. PCC 6803 as a frequent modification of proteins associated with photosynthesis and enzymes in the central carbon metabolism1. Strong abundance changes of specific phosphorylation events were detected by quantitative phosphoproteomics in response to the autotrophy-to-heterotrophy switch during resuscitation from nitrogen starvation. Regulatory functions of individual phosphorylation events have recently been demonstrated2; however, the function of most of them is so far unknown.
In the SCyCode consortium, we teamed up with the leading research groups in cyanobacterial metabolism control to characterize the post-translational mechanisms involved in cellular regulation and survival during autotrophy-to-heterotrophy switches. So far, together with the Forchhammer group we could show that phosphorylation is a key to control enzyme activity in glycogen catabolism3, and that specific protein-protein interactions regulate glycogen accumulation4 and phycobiliprotein degradation5.
Furthermore, together with Hagemann and Hess groups we characterized the substrates of protein kinases relevant in cellular acclimation to carbon limitation6, RNA-protein and protein-protein interactions by gradient profiling (GradSeq)7 and extended this strategy to the detection of protein-metabolite interactions. Finally, we developed a proteogenomics pipeline to screen our large-scale proteomics data for non-annotated protein coding genes and thus to provide a re-annotated map of the Synechocystis genome for the entire research community.
1 Spät, P., Maček, B. & Forchhammer, K. (2015) Phosphoproteome of the cyanobacterium Synechocystis sp. PCC 6803 and its dynamics during nitrogen starvation. Frontiers in Microbiology 6, doi:10.3389/fmicb.2015.00248
2 Spät, P., Klotz, A., Rexroth, S., Maček, B. & Forchhammer, K. (2018) Chlorosis as a Developmental Program in Cyanobacteria: The Proteomic Fundament for Survival and Awakening. Molecular & cellular proteomics: MCP 17, 1650-1669, doi:10.1074/mcp.RA118.000699
3 Doello, S., Neumann, N., Spät, P., Maček, B & Forchhammer, K. Phosphoglucomutase as central hub in the control of glycogen metabolism (submitted)
4 Orthwein, T, Scholl, J., Spät, P., Lucius, S., Koch, M., Macek, B. Hagemann, M. & Forchhammer, K. (2021) The novel PII-interactor PirC identifies phosphoglycerate mutase as key control point of carbon storage metabolism in cyanobacteria. PNAS e2019988118
5 Krauspe, V., Fahrner, M., Spät, P., Steglich, C., Frankenberg-Dinkel, N., Maček, B., Schilling, O., and Hess, W.R. (2021). Discovery of a small protein factor involved in the coordinated degradation of phycobilisomes in cyanobacteria. Proceedings of the National Academy of Sciences 118, e2012277118.
6 Spät, P., Barske T, Maček, B., and Hagemann, M. Alterations in the CO2 availability induce alterations in the phospho-proteome of the cyanobacterium Synechocystis sp. PCC 6803. New Phytol (accepted)
7 Riediger, M., Spät, P., Bilger, R., Voigt, K., Maček, B., and Hess, W.R. (2020). Analysis of a Photosynthetic Cyanobacterium Rich in Internal Membrane Systems via Gradient Profiling by Sequencing (Grad-seq). The Plant Cell, doi:10.1093/plcell/koaa017
Principal Investigator

University of Tübingen
Department of Quantitative Proteomics
Auf der Morgenstelle 15, 72076 Tübingen, Germany
boris.macekuni-tuebingen.de
Website
Members

niels.neumannuni-tuebingen.de

philipp.spaetuni-tuebingen.de

Alumni

Alumni